Contiguous Gene Deletion of Chromosome Xp in Three Families Encompassing OTC, RPGR and TSPAN7 Genes
Shailly Jain-Ghai,Stephanie Skinner, Jessica Hartley,Stephanie Fox, Daniela Buhas, Cheryl Rockman-Greenberg and Alicia Chan
1Medical Genetics Clinic, Stollery Children's Hospital, Edmonton, Alberta,Canada
2Program in Genetics and Metabolism,Winnipeg Regional Health Authority and University of Manitoba, Winnipeg,Manitoba, Canada
3Department of Biochemistry and Medical Genetics, University of Manitoba, Winnipeg, Manitoba, Canada
4Department of Medical Genetics,Montreal Children’s Hospital, McGill University Health Centre, Montreal,Quebec, Canada
5Department of Medical Genetics,University of Alberta, Edmonton,Alberta, Canada
- *Corresponding Author:
- Shailly Jain
Medical Genetics Clinic, 8-53 Medical Sciences Building, University of Alberta Hospital
dmonton, AB, Canada T6G 2H7
Tel: 1-(780) 407-7333
E-mail: Shailly.jain@albertahealthservices.ca
Citation: Ghai SJ, Skinner S, et al. Contiguous Gene Deletion of Chromosome Xp in Three Families Encompassing OTC, RPGR and TSPAN7 Genes. J Rare Dis Diagn Ther. 2015, 1:1. doi:10.21767/2380-7245.10003
Copyright: © 2015 Ghai SJ, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Ornithine transcarbamylase deficiency (OTCD) is the most common urea cycle disorder. The classic presentation in males is hyperammonemic encephalopathy in the early neonatal period. Given the X-linked inheritance of OTCD, presentation in females is highly variable. We present three families with different contiguous gene deletions on chromosome Xp. Deletion of RPGR, OTC and TSPAN7 is common to all three families in our series. These cases highlight the variable phenotype in manifesting OTCD female carriers, the complexity of OTCD management and complex issues surrounding the option of liver transplantation when multiple other genetic factors play a role.
Keywords
Ornithine transcarbamylase; Ornithine transcarbamylase deficiency; Contiguous gene deletion; OTC;RPGR; TSPAN7
Abbreviations
CGH: Comparative Genomic Hybridization; CRRT: Continuous Renal Replacement Therapy; MLPA: Multiplex Ligation-Dependent Probe Amplification; OTCD: Ornithine Transcarbamylase Deficiency; OTC: Ornithine Transcarbamylase SNP: Single Nucleotide Polymorphism
Introduction
The urea cycle functions to convert ammonia, a byproduct of protein breakdown, into urea, which can be safely excreted by the body. The most common urea cycle disorder, ornithine transcarbamylase deficiency (OMIM 311250) (OTCD), is an X-linked condition with an incidence of 1:14000 [1]. The majority of males with OTCD present with early neonatal hyperammonemic encephalopathy which predisposes to neurological insults and early death. Use of renal hemodialysis or equivalent, ammonia scavenger medications, synthetic formula and dietary protein restriction can allow for survival into infancy, at which point liver transplantation can be considered. For heterozygous female carriers, an estimated 20% have symptoms, ranging from neonatal hyperammonemic encephalopathy to adult onset behavioral changes and cognitive delays [2,3].
The combination of hyperammonemia, elevated plasma glutamine and elevated urinary orotic acid in addition to a possible decrease in plasma citrulline leads to suspicion of OTCD. Liver enzyme assay can be performed for diagnostic confirmation; however it is technically difficult. Thus, confirmation is most often achieved by molecular analysis; and is considered the gold standard of testing in females and in prenatal cases [4]. Mutations are detected in only about 80% of all OTC cases [5]. Of these, over 80% are missense or nonsense mutations [6] and 10-15% may be due to partial or complete gene deletions [7]. With the use of array CGH, MLPA and high density SNP array, many contiguous gene deletions involving the OTC gene are being described [4, 8-13].
We describe three unrelated families with different contiguous gene deletions on chromosome Xp. Deletion of RPGR, OTC and TSPAN7 is common to all these families. We highlight the impact of deletions of multiple genes surrounding the OTC locus on the ultimate OTCD phenotype and its management.
Clinical Report
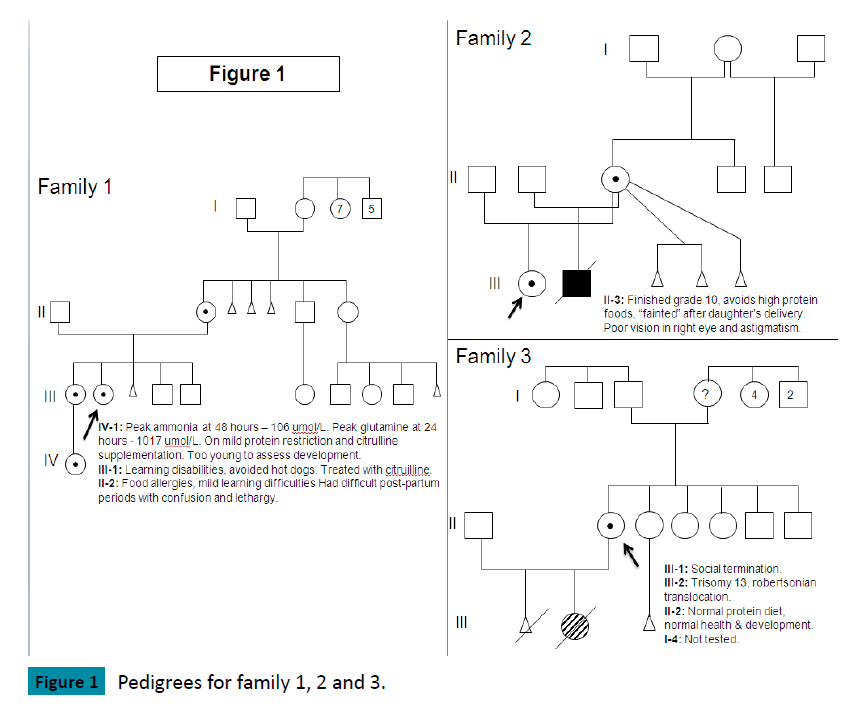
Family 1 (Figure 1)
The index case, female, presented at 12 years of age with delirium, emesis, weakness and weight loss. Investigations at diagnosis showed: ammonia 242 umol/L (normal 5-35 umol/L), glutamine 2000 umol/L (normal: 450-750), citrulline 25 umol/L (1-40), ornithine 103 umol/L (50-100) but orotic acid was not elevated. Past medical history revealed developmental delay noted at 8-12 months of age. She had a tendency towards recurrent vomiting, headaches, lethargy and abnormal behavior, which improved with avoidance of meat and dairy products. She had mild congenital optic nerve hypoplasia. At age 12, she was functioning at the level of a 3-4 year old. MRI brain showed cortical atrophy in left frontal region. Since diagnosis, treatment has consisted of dietary protein restriction supplemented with cyclinex formula, sodium phenylbutyrate and L-citrulline. She is currently 24 years old, has cognitive delays, lacks executive functioning and is unable to live independently. Since diagnosis, her highest ammonia was 406 umol/L but glutamine never exceeded 2000 umol/L. She has had multiple admissions due to hyperammonemic encephalopathy but does not have other medical complications.
Her older sister had learning difficulties and was “grouchy” during her menstrual periods. She avoided hot dogs which caused headache and vomiting but tolerated (other) high protein foods. Her investigations at diagnosis showed: ammonia 27 umol/L, glutamine 1100 umol/L, citrulline 6 umol/L, ornithine 125 umol/L. L-Citrulline supplementation has been her only treatment. Since diagnosis, her peak ammonia was 95 umol/L and glutamine never exceeded 1100 umol/L. She recently had an uneventful pregnancy and delivery, with peak post partum ammonia of 64 umol/L and glutamine of 852 umol/L.
Family 2 (Figure 1)
The index case, female, presented at 7 months of age with back arching and an abnormal EEG. At diagnosis, ammonia was 388 umol/L, ornithine 32 umol/L, citrulline undetectable, glutamine 1112 umol/L with an elevated orotic acid. Past medical history showed recurrent vomiting episodes starting at two weeks of age, failure to thrive and developmental delay. MRI brain showed abnormal myelination with microcephaly and atrophy. At 10 years of age, she has been seizure free but has significant developmental delays. She also has nystagmus, amblyopia, myopia and bilateral thin optic nerves. Her current treatment consists of protein restriction supplemented with Cyclinex formula, L-citrulline and sodium phenylbutyrate.
In a subsequent pregnancy, her mother declined prenatal diagnosis and gave birth to a boy. His ammonia steadily increased and was 512 umol/L by day 2 of life, requiring CRRT (dialysis). He was later maintained on a protein restricted diet, L-citrulline, sodium phenylbutyrate and Cyclinex formula. He developed failure to thrive by 4 months of age as trials of increasing dietary protein led to hyperammonemia. At 12 months, he presented with liver disease (hypoalbuminemia, edema, ascites, and mild coagulopathy), hypothyroidism, recurrent need for pRBC transfusion, growth failure, developmental delay (functioned at a 2-3 month level) and skin rashes. At this time, trials of higher protein (1 – 1.2 grams of protein/kg/day) were tolerated without occurrence of hyperammonemia but normalization of protein status did not lead to improvement in his overall clinical picture. Ammonia levels did not peak higher than the level at diagnosis. He became ventilation dependent and was not considered to be a candidate for liver transplant. He died due to multi-organ failure at 14 months of age.
Family 3 (Figure 1)
The index case, a 24 year old female with no health issues or developmental concerns, gave birth to a girl. This baby had Trisomy 13 due to a Robertsonian translocation. Cytogenetic investigations of the index case and her partner were normal with respect to the translocation but incidentally identified an X-chromosome deletion for the index case. At diagnosis, her ammonia was normal, with glutamine 487 umol/L, citrulline 19 umol/L, and arginine 74 umol/L. She did not restrict dietary protein and had no developmental or cognitive concerns. She was studying psychology in University.
Genetic Testing
Family 1: Molecular analysis for OTC was not available at the time of initial diagnosis. A karyotype on the index case showed an estimated 5Mb deletion of Xp21.1 (46,X,del(X)(p21.1p21.1)) at 600 band resolution. To refine the breakpoints, array CGH was ordered in 2012. Testing identified a heterozygous copy number loss of 8.05 Mb on Xp between nucleotides 30,376,093-38,432,866 (NCBI36/hg18; CytoChip ISCA 8x60K v2.0), corresponding to a cytogenetic location of Xp11.4-Xp21.2. This correlates with a loss of 25 RefSeq protein coding genes.
Family 2: Sequencing of the OTC gene on the index case repeatedly failed to amplify any gene product, suggesting a gross deletion. An array CGH on the brother of the index case identified a hemizygous loss at Xp11.4 with a minimal interval spanning from nucleotides 38,068,450-38,810,601, corresponding to a loss of 0.742 Mb (GRCh37/hg19, Oligo V8.1.1). This correlates with a loss of 5 RefSeq protein-coding genes.
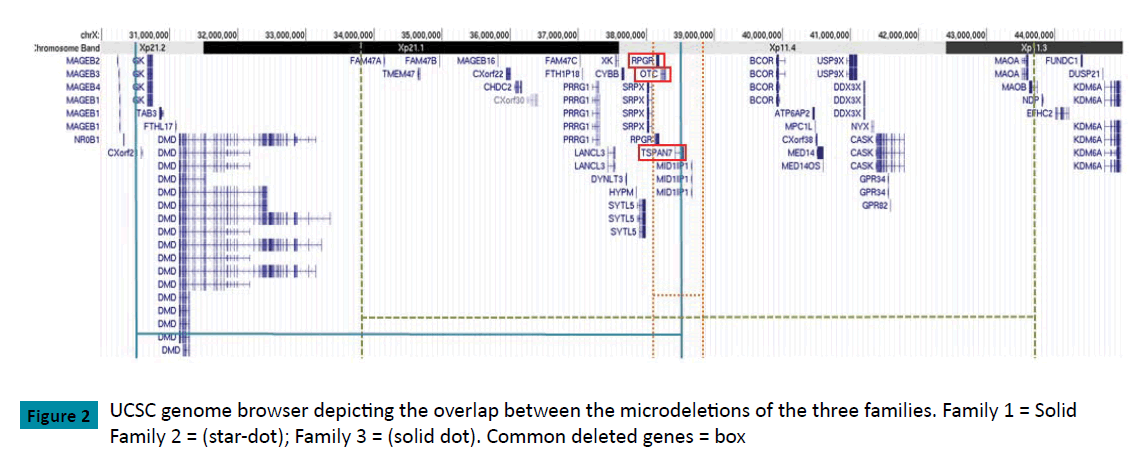
Family 3: Following the finding of abnormal karyotype, array CGH on the index case confirmed a heterozygous copy number loss of 9.88 Mb between nucleotides 33,798,269-43,675,842 corresponding to a cytogenetic location of Xp11.3-Xp21.1 (GRCh37/hg19). This correlates with a loss of 35 Refseq proteincoding genes.(Figure 2,Table 1)
| Gene | Family 1 | Family 2 | Family 3 | Phenotype (OMIM #) | |
|---|---|---|---|---|---|
| Karyotype Coordinates | Xp11.4-Xp21.2 | Xp11.4 | Xp11.3-Xp21.1 | ||
| Estimated Deletion Breakpoints (hg19) | chrX:30,466,172-38,547,922* | chrX:38,068,450-38,810,601 | chrX:33,798,269-43,675,842 | ||
| GK | Glycerol kinase deficiency (300474) | ||||
| DMD | Duchenne muscular dystrophy; Becker muscular dystrophy; dilated cardiomyopathy (300377) | ||||
| XK | McLeod syndrome, with or without chronic granulomatous disease (314850) | ||||
| CYBB | Familial atypical mycobacteriosis / Chronic granulomatous disease (300481) | ||||
| RPGR | Cone-rod dystrophy; Macular degeneration; Retinitis pigmentosa; Retinitis pigmentosa, X-linked, and sinorespiratory infections, with or without deafness (312610) | ||||
| OTC | Ornithine transcarbamylase deficiency (300461) | ||||
| TSPAN7 | X-linked mental retardation (300096) | ||||
| BCOR | Syndromicmicrophthalmia; oculo-facio-cardio-dental syndrome (300166) | ||||
| ATP6AP2 | Mental retardation X-linked, Hedera type (300423) | ||||
| USP9X | Mental retardation, X-linked 99 (300919) | ||||
| NYX | Night blindness, congenital stationary (complete), 1A, X-linked (310500 ) | ||||
| CASK | FG syndrome 4 (300422); Mental retardation, with or without nystagmus (300422); Mental retardation and microcephaly with pontine and cerebellar hypoplasia (300749) | ||||
| MAOA | Brunner syndrome (300615) |
Black denotes complete deletion, hatching denotes partial deletion.
Table 1: Contiguous gene deletions and OMIM genes for the three families.
Discussion
The X chromosome has been noted to be a hot spot for genomic rearrangement, and is thus predisposed to copy number changes such as deletions [14]. Plausible explanations for Xp region copy number changes include intramolecular homologous recombination [15, 16] and/or DNA slippage [16]. With the current technology of CGH and high density SNP array, it has become possible to identify microdeletions that may have previously been missed by karyotype, and it has also become much easier to identify the extent of deletions, both large and small. This allows us to better understand the phenotypic alterations in metabolic conditions like OTCD.
The cases presented here highlight the variable clinical features in males and females with contiguous gene deletions involving the OTC gene. The two index cases from Family 1 and 2 had biochemical features of OTCD at presentation but molecular and cytogenetic studies revealed a more extended chromosomal microdeletion rather than the initially suspected monogenic disorder. In contrast, the index case of Family 3 was identified incidentally without any clinical or biochemical features of OTCD. The severity of OTCD in carrier females is certainly related to the degree of X-chromosome inactivation but it remains unclear if other genes modify the OTCD phenotype. In these families, their clinical presentation is further complicated by the extent of the deletion, as it can become difficult to tease apart the cognitive symptoms due to OTCD from those related to deletion of the surrounding genes. Deardroff et al. [11] have commented on a number of individuals with deletions of Xp11.4-Xp21.2 with intact OTC gene(s) that had minimal, if any, cognitive delays [11]. Similar to our experience with Family 1, Balasubramaniam et al. [4] report a female with contiguous gene deletion of the OTC, DMD (Duchenne muscular dystrophy), RPGR (cone-rod dystrophy/ retinitis pigmentosa), CYBB (chronic granulomatosis disease) and XK (McLeod syndrome) genes, manifesting only symptoms of OTCD and mild developmental delay. In our case series, carrier females in Family 1 have not developed symptoms of other conditions like Duchenne muscular dystrophy (DMD) (normal CK and echocardiograms), glycerol kinase (GK) (normal urine organic acids), chronic granulomatous disease (CYBB) (absent clinical symptoms) and McLeod syndrome (XK) (normal blood smear); however, carrier females in both Family 1 and Family 2 manifest symptoms which may be related to RPGR and TSPAN7 , which are associated with ophthalmic disease and X-linked mental retardation respectively. Index females in both families have hyperammonemia, significant ophthalmological findings and a severity of developmental delay that may not be fully explained by only hyperammonemia. Even in the milder phenotypes of other carrier females in these two families, learning difficulties and mild vision concerns are prominent in absence of documented hyperammonemia (Figure 1), but in the presence of mild OTCD symptoms such as protein avoidance or biochemical disturbances. Taken together, there appears to be an all or none phenomenon with respect to phenotypic expression. The families seem to either present with symptoms of all three conditions or remain asymptomatic with respect to all three, which is likely explained by the proximity of these genes to each other. Thus, Family 1 and 2 demonstrate symptoms of all three genes, with consistency of severity within an individual compared to Family 3 where symptoms of none of these three deleted genes are present.
As males are hemizygous, contribution of all deleted genes to the phenotype is expected and may allow for an explanation when routine OTCD management is not effective. In the male case presented here (Family 2), ascites and anasarca was initially felt to be due to severe protein restriction as a result of OTCD treatment. However, even with correction of protein status, anasarca did not resolve and the baby progressed to having respiratory complications. The RPGR deletion and its association with primary cilliary dyskinesia [17] and sinorespiratory infections (OMIM 300455) likely contributed to his clinical picture. RPGR is a cilia-centrosomal protein believed to cause cilia-dependent photoreceptor degeneration [18] but may also play a role in pulmonary cilia orientation [19]. Other reported systemic features have included hearing loss [20]. Deardroff et al [11] presented a similar scenario where a male patient with deletion of RPGR and OTC developed anasarca at 3 weeks of age despite having had good metabolic control. Given the close proximity of RPGR to OTC, it is important to exclude its deletion in cases where unexpected complications of OTCD are seen. Liver transplant, an acceptable treatment for males with severe OTCD, may not be suitable if RPGR is also deleted as it will increase the likelihood of respiratory complications pre- and post-transplant. Discussion about the possibility of adult-onset blindness in these cases may also be warranted. Little is known about TSPAN7 except for its implications in non-syndromic non-specific X-linked mental retardation (MR) [21,22] possibly due to dys-regulation at neuronal glutamatergic synapses [23]. Phenotypically, patients are described to have mild to moderate MR [24] and thus deletion of TSPAN7 can be an additional risk factor for poor intellectual development independent of OTCD.
Families 1 and 2 have both reported multiple miscarriages and this raises the possibility of prenatal lethality, likely in males, due to the microdeletion. This again emphasizes that copy number aberrations of this gene rich region severely hamper development of multiple organ systems.
In conclusion, we present three families with Xp contiguous deletion of the region encompassing OTC, RPGR and TSPAN7 with multiple members, all displaying variable phenotypes. Cases like these highlight the need to fully elucidate the underlying molecular abnormality as it will alter treatment and possibility of candidacy for liver transplant in OTCD. It also allows us to better understand function of surrounding genes and their possible interactions.
Conflict of Interest
The authors have no conflict of interest to disclose
Acknowledgement
We acknowledge the families and patients for providing consent and their support for this project.
References
- Batshaw ML, Lichter-Konecki U, Tuchman M (2006) Inborn errors of urea synthesis. In: Swaiman KF, Ashwal S, Ferriero DM (eds) Pediatric Neurology: Principles & Practice. (4thedn), Mosby Elsevier, USA.
- Arn PH, Hauser ER, Thomas GH, Herman G, Hess D, et al. (1990) Hyperammonemia in women with a mutation at the ornithine carbamoyltransferase locus, a cause of postpartum coma. N Engl J Med 322: 1652–1655.
- Nicolaides P, Liebsch D, Dale N, Leonard J, Surtees R (2002) Neurological outcome of patients with ornithine carbamoyltransferase deficiency. Arch Dis Child 86: 54–56.
- Balasubramaniam S, Rudduck C, Bennetts B, Peters G, Wilcken B, et al. (2010) Contiguous gene deletion syndrome in a female with ornithine transcarbamylase deficiency. Molec Genet Metab 99: 34–41.
- Tuchman M, Jaleel N, Morizono H, Sheehy L, Lynch MG (2002) Mutations and polymorphisms in the human rnithine transcarbamylase (OTC) gene. Hum Mutat 19: 93–107.
- Yamaguchi S, Brailey LL, Morizono H, Bale AE, Tuchman M (2006) Mutations and polymorphisms in the human rnithine transcarbamylase (OTC) gene. Hum Mutat 27: 626–632.
- Tuchman M (1993) Mutations and polymorphisms in the human ornithine transcarbamylase gene. Hum Mutat 2: 174–178.
- Ono M, Tsuda J, Mouri Y, Arai J, Arinami T, et al. (2010) Contiguous Xp11.4 Gene Deletion Leading to Ornithine ranscarbamylase Deficiency Detected .
- Storkanova G, Vlaskova H, Chuzhanova N, Zeman J, Stranecky V, et al. (2013) Ornithine carbamoyltransferase deficiency: molecular characterization of 29 families. Clin Genet 84: 552–559.
- Quental R, Azevedo L, Rubio V, Diogo L, Amorim A (2009) Molecular mechanisms underlying large genomic deletions in ornithine transcarbamylase (OTC) gene. Clin Genet 75: 457–464.
- Deardroff MA, Gaddipati H, Kaplan P, Sanchez-Lara PA, Sondheimer N, et al. (2008) Complex management of a patient with a contiguous Xp11.4 gene deletion involving ornithine transcarbamylase: A role for detailed molecular analysis in complex presentations of classical diseases. Molec Genet Metab 94: 498-502.
- Arranz JA, Madrigal I, Riudor E, Armengol L, Milà M (2007) Complete deletion of ornithine transcarbamylase gene confirmed by CGH array of X chromosome. J Inherit Metab Dis 30: 813.
- Jakubiczka S, Bettecken T, Mohnike K, Schneppenheim R, Stumm M, et al. (2007) Symptoms of OTC deficiency but not DMD in a female carrier of an Xp21.1 deletion including the genes for dystrophin and OTC. Eur J Pediatr 166: 743–745.
- Todorova A, Litvinenko I, Todorov T, Tincheva R, Avdjieva D, et al. (2014) A family with fragile X syndrome, uchenne muscular dystrophy and ichthyosis transmitted by an asymptomatic carrier. Clin Genet 85: 286–289.
- Xiao-Miao L, Yen PH, Shapiro LJ (1992) Characterization of a low copy repetitive element S232 involved in the generation of frequent deletions of the distal short arm of the human X chromosome. Nucleic Acids Res 20: 1117–1122.
- Cuevas-Covarrubias SA, Jiménez-Vaca AL, González-Huerta LM, Valdes-Flores M, Kofman-Alfaro SH, et al. (2002) Somatic and germinal mosaicism for the steroid sulfatase gene deletion in a steroid sulfatase deficiency carrier. J Invest Dermatol 119: 972 – 975.
- Moore A, Escudier E, Roger G, Tamalet A, Pelosse B, et al. (2006) RPGR is mutated in patients with a complex X linked phenotype combining primary ciliary dyskinesia and retinitis pigmentosa. J Med Genet 43: 26–33.
- Murga-Zamalloa CA, Swaroop A, Khanna H (2009) RPGR-containing protein complexes in syndromic and nonsyndromic retinal degeneration due to ciliary dysfunction. J Genet 88: 399–407.
- Bukowy-Bieryłło Z, Ziętkiewicz E, Loges NT, Wittmer M, Geremek M, et al. (2013) RPGR Mutations Might Cause Reduced Orientation of Respiratory Cilia. Pediatr Pulmonol 48: 352–363.
- Zito I, Downes SM, Patel RJ, Cheetham ME, Ebenezer ND, et al. (2003) RPGR mutation associated with retinitis pigmentosa, impaired hearing, and sinorespiratory infections. J Med Genet 40: 609-615.
- Abidi FE, Holinski-Feder E, Rittinger O, Kooy F, Lubs HA, et al. (2002) A novel 2 bp deletion in the TM4SF2 gene is associated with MRX58. J Med Genet 39: 430-433.
- Zemni R, Bienvenu T, Vinet MC, Sefiani A, Carrié A, et al. (2000) A new gene involved in X-linked mental retardation identified by analysis of an X;2 balanced translocation. Nature Genet 24: 167-170.
- Bassani S, Cingolani LA, Valnegri, Folci A, Zapata J, et al (2012) The X-linked intellectual disability protein TSPAN7 regulates excitatory synapse development and AMPAR trafficking. Neuron 73: 1143-1158.
- Holinski-Feder E, Chahrockh-Zadeh S, Rittinger O, Jedele KB, Gasteiger M, et al. (1999) Nonsyndromic X-linked mental retardation: mapping of MRX58 to the pericentromeric region. Am J Med Genet 86: 102-106.
Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences